The Encyclopedia of DNA Elements (ENCODE) Consortium is an international collaboration of research groups funded by the National Human Genome Research Institute (NHGRI). The goal of ENCODE is to build a comprehensive parts list of functional elements in the human genome, including elements that act at the protein and RNA levels, and regulatory elements that control cells and circumstances in which a gene is active.
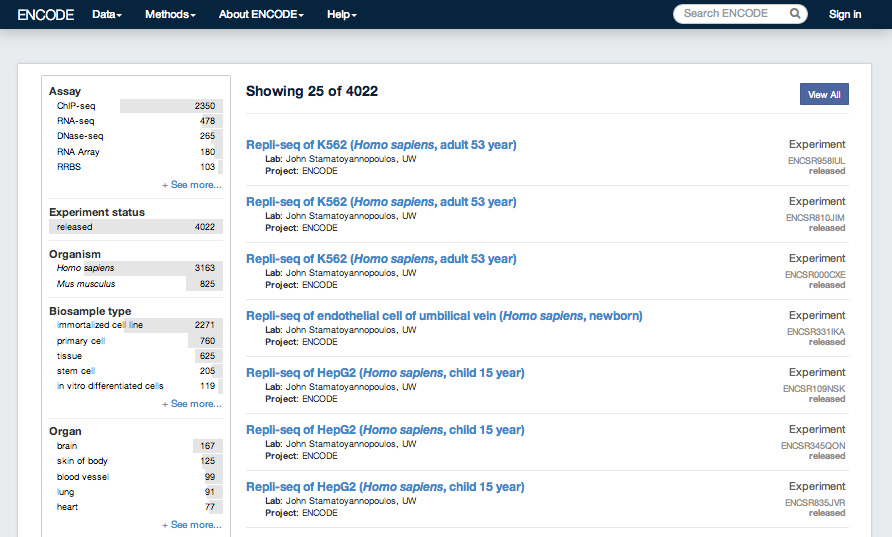
ENCODE results from 2007 and later are available from the ENCODE Project Portal, encodeproject.org. This covers data generated during the two production phases 2007-2012 and 2013-present. The ENCODE Project Portal also hosts additional ENCODE access tools, and ENCODE project pages including up-to-date information about data releases, publications, and upcoming tutorials.
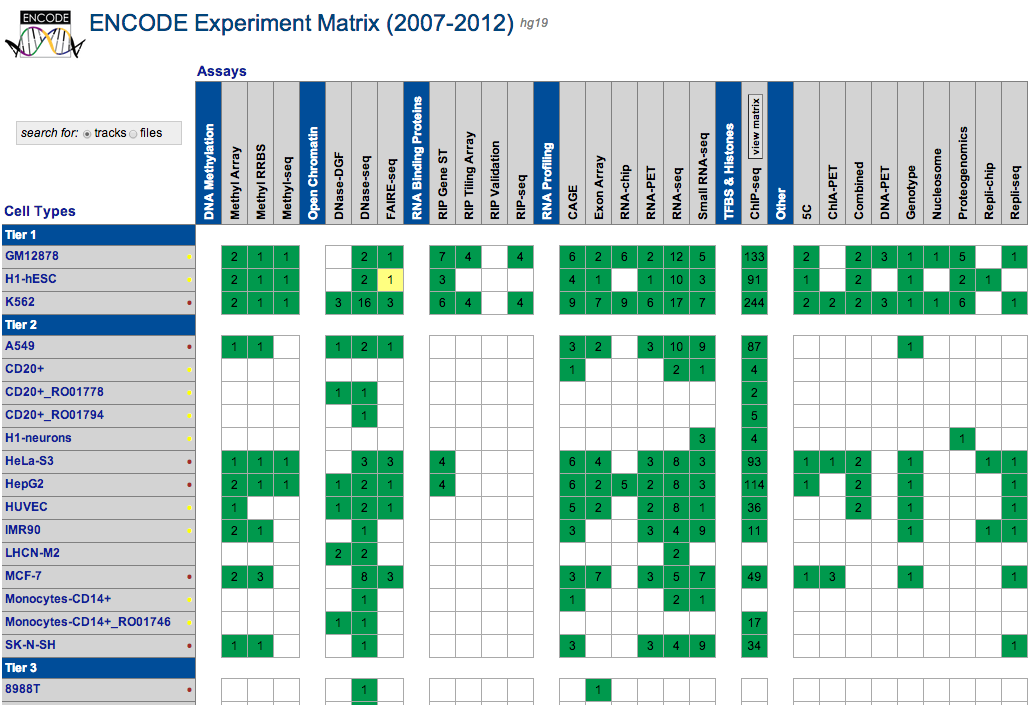
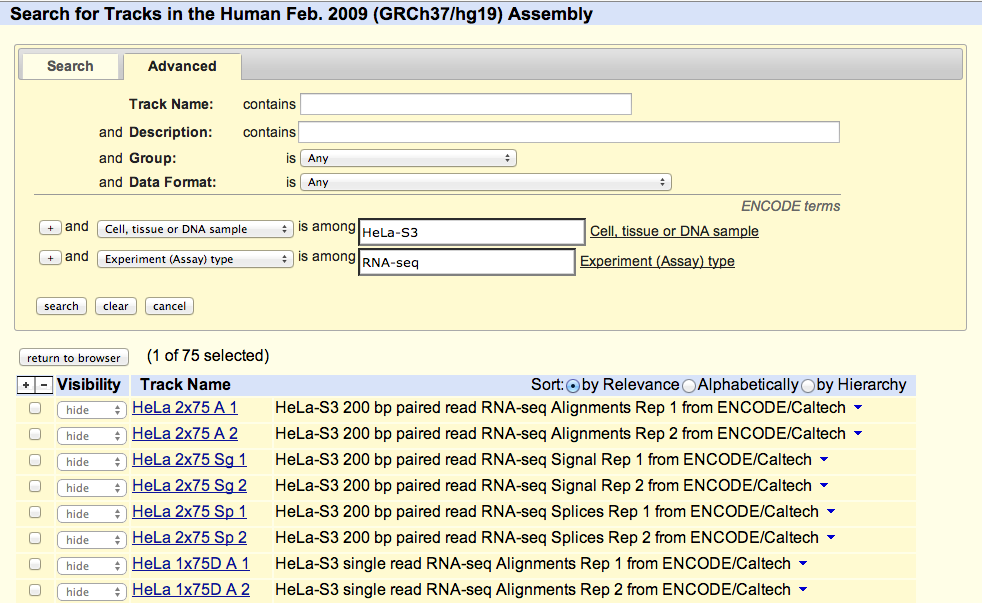
UCSC coordinated data for the ENCODE Consortium from its inception in 2003 (Pilot phase) to the end of the first 5 year phase of whole-genome data production in 2012. All data produced by ENCODE investigators and the results of ENCODE analysis projects from this period are hosted in the UCSC Genome browser and database. Explore ENCODE data using the image links below or via the left menu bar. All ENCODE data at UCSC are freely available for download and analysis.